
Recent Posts:
High Throughput Insights Into Microbial Metabolism: SymproveTM Modulates Gut Microbiome Function Across Healthy and Dysbiotic States
Understanding how probiotics influence the human gut microbiome requires tools capable of resolving not only community composition, but also the full breadth of metabolic activity emerging from microbe-microbe and microbe-substrate interactions. In the current study, we have applied our Colon‑on‑a‑Plate® (CoaP) model in combination with Laser‑Assisted Rapid Evaporative Ionization Mass Spectrometry (LA‑REIMS), targeted Short-Chain Fatty Acid (SCFA) analysis and in-depth metabolomics using Ultra-High-Performance Liquid Chromatography coupled with High-Resolution Mass Spectrometry (UHPLC‑HRMS) to investigate how the multi-strain probiotic SymproveTM affects microbial metabolism under a wide array of physiologically relevant conditions. These conditions included a healthy microbiome, microbiomes with antibiotic‑induced dysbiosis (three severity levels, across five antibiotic classes), and environments challenged with enterotoxigenic Escherichia coli (ETEC).
The study provides one of the most comprehensive datasets to date on probiotic-induced metabolic changes, demonstrating consistent effects across hundreds of experimental conditions. Importantly, it reveals mechanistic insights that extend beyond classical SCFA profiling, highlighting biochemical pathways relevant to intestinal health, immune signaling and even the gut-brain axis.
Experimental Design and Analytical Framework
Fecal samples from 10 healthy adult donors were used to establish parallel ex vivo colonic microbiome simulations. A single dose of SymproveTM (containing Lacticaseibacillus acidophilus, Lactiplantibacillus plantarum, Lacticaseibacillus rhamnosus and, Enterococcus faecium) was added to our CoaP system and compared to untreated controls. To systematically explore microbial responses, the team employed a fractional factorial design, generating 576 distinct test conditions. Microbial metabolism was assessed at 24h and 48h, combining:
- LA‑REIMS for untargeted metabolic fingerprinting
- Targeted SCFA and branched SCFA quantification
- UHPLC‑HRMS for deep metabolic profiling of >400 polar to medium polar metabolites
This approach enabled detailed pathway‑level interpretation of microbial responses with high statistical power, despite extensive condition diversity.
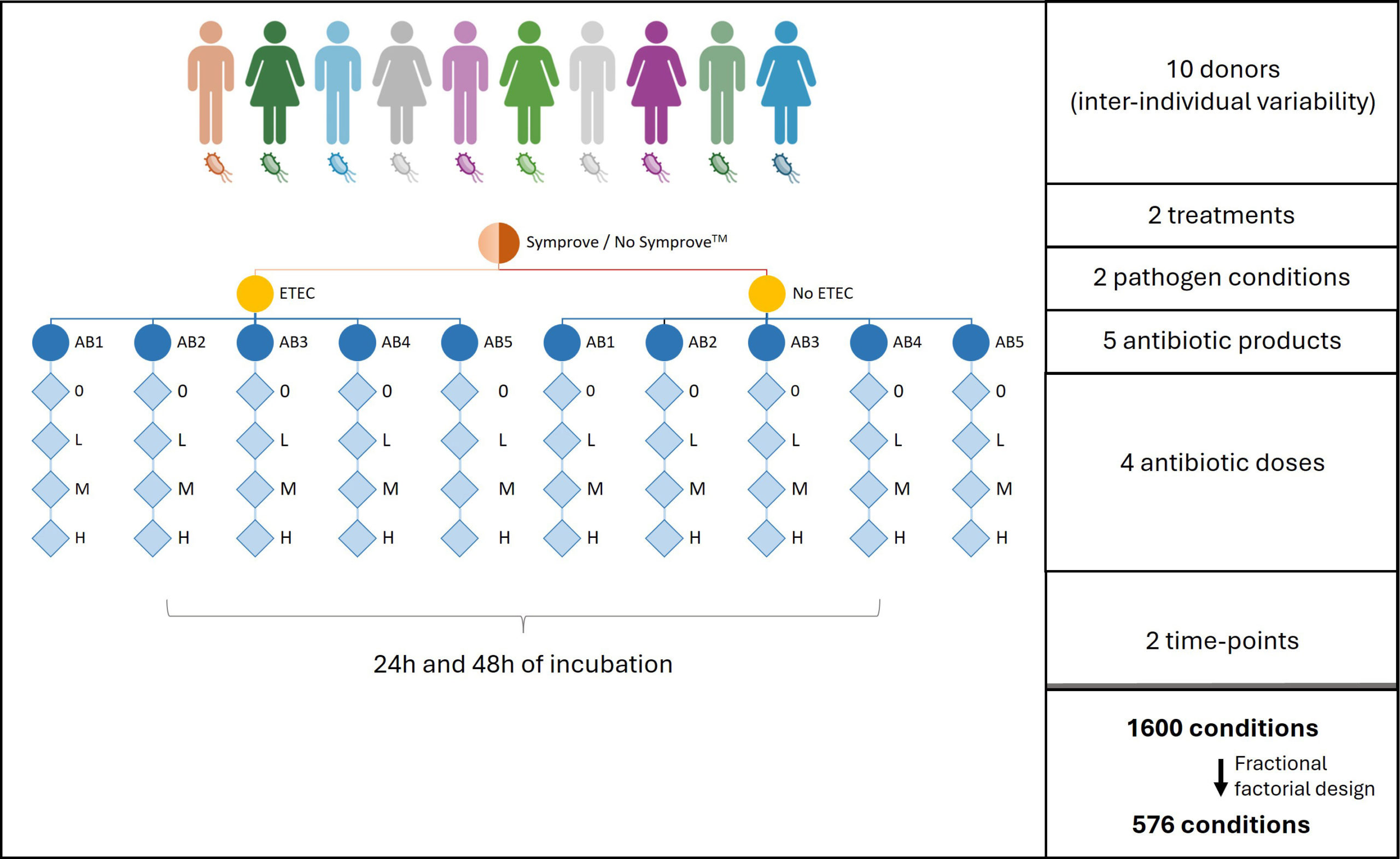
Study design. Individual fecal samples were collected from 10 healthy donors and used in CoaP platform with or without antibiotic pre-incubation (one of five antibiotics at a low, medium, or high concentration). Samples were either infected with ETEC or control media (no ETEC). Under each described condition, samples were supplemented with Symprove™ or control media (no Symprove™). Finally, each condition was set up twice, to allow for evaluation at two time points (24 h or 48 h). 0: no antibiotic addition; AB1-5: antibiotic 1-5; ETEC: enterotoxigenic Escherichia coli; H: high antibiotic dose; L: low antibiotic dose; M: medium antibiotic dose.
Effects on SCFA Production and Fermentation Balance
Initial untargeted LA‑REIMS fingerprinting revealed that Symprove™ shifted overall microbial metabolism for both healthy and dysbiotic microbiomes. These changes were already visible at 24 h (as illustrated in the figure below) and remained present at 48 h. Multivariate modelling confirmed that these shifts were statistically robust.
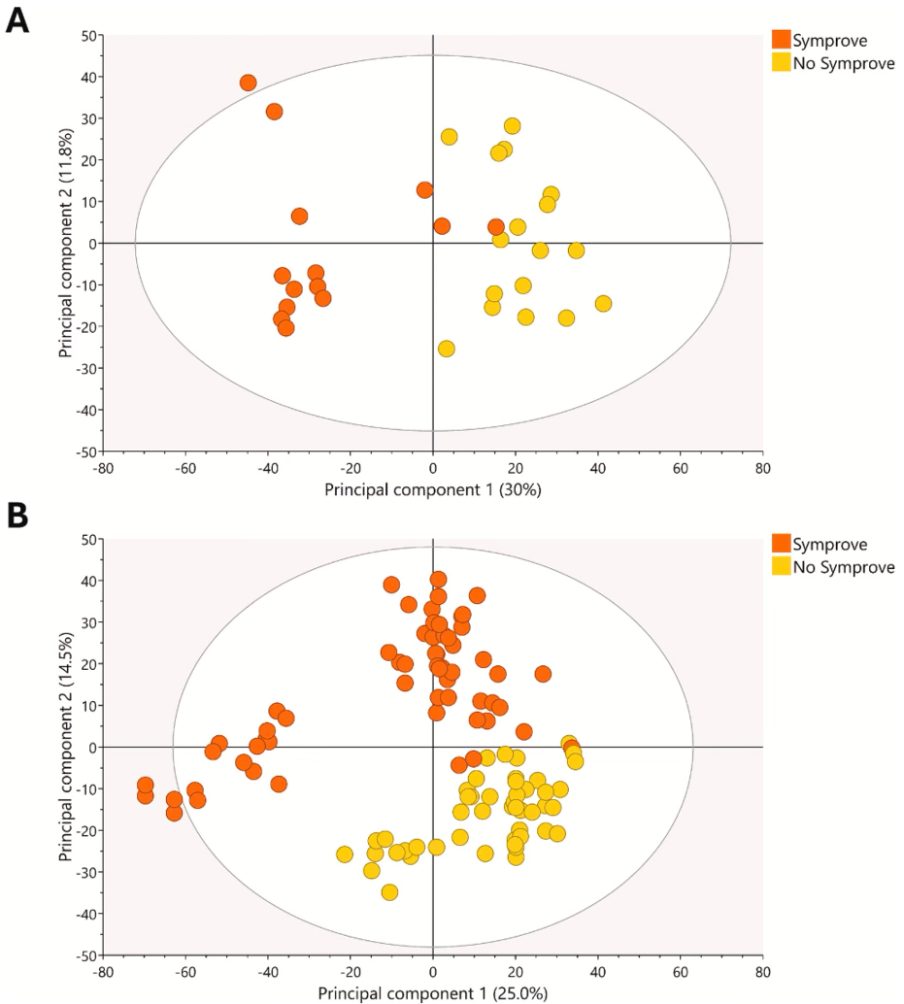
PCA-X plots based on the LA-REIMS data for the healthy population (no ETEC infection and no antibiotic use) (A) and for the dysbiosed population (no ETEC infection and following antibiotic use) (B). LA-REIMS was used to measure the metabolic activity of colonic fermentations collected after 24 h incubation with Symprove™ or untreated control (water) in the Colon-a-Plate® model using fecal samples from 10 healthy human donors. ETEC: enterotoxigenic Escherichia coli; LA-REIMS: Laser-Assisted Rapid Evaporative Ionization Mass Spectrometry; OPLS-DA: orthogonal partial least squares discriminant analysis; PCA-X: unsupervised principal component analysis.
Targeted SCFA analysis showed that across both healthy and dysbiotic microbiomes, SymproveTM consistently enhanced saccharolytic fermentation, illustrated by significant increases in acetate, propionate and butyrate. For example, in healthy microbiota at 24h, acetate and propionate significantly increased by about 1.5-fold, whereas this was with more than 2-fold for butyrate. These effects persisted under increasing dysbiosis severity, suggesting the probiotic retains functional metabolic impact even when community structure is compromised.
Simultaneously, branched SCFAs (markers of protein fermentation) were significantly reduced across all conditions. Because excessive proteolytic fermentation is associated with inflammation and epithelial stress, this shift indicates a more metabolically favorable gut environment promoted by SymproveTM. The same SCFA and branched SCFA patterns were observed when dysbiotic microbiomes were further challenged with ETEC, underscoring that Symprove™ can restore healthier fermentation profiles even in severely disrupted environments.
Anti‑inflammatory and Gut Barrier‑Supporting Pathways
Beyond SCFAs, SymproveTM altered numerous metabolites with known roles in immune regulation and epithelial gut barrier maintenance, supporting previously reported effects of SymproveTM.
Key observations obtained via in-depth metabolomics UHPLC‑HRMS include:
- Elevated levels of hydroxy fatty acids (e.g. 3‑hydroxybutyric acid) which exhibit anti-inflammatory and immunomodulatory properties.
- Increases in indole‑3‑lactic acid and indole‑3‑acetic acid, both derived from probiotic tryptophan metabolism and known to strengthen mucosal immunity.
- Enhanced purine and pyrimidine metabolites (e.g. inosine, uracil, cytidine) potentially contributing to cellular repair processes and mucosal regeneration.
- Increased amino‑acid acetylation possibly reflecting shifts in host-microbe protein modification pathways relevant to the intestinal barrier.
Gut-Brain Axis Signatures: GABA and L‑DOPA Modulation
A compelling aspect of our study is the modulation of metabolites linked to the gut-brain axis. SymproveTM supplementation increased γ‑aminobutyric acid (GABA) levels across both healthy and dysbiotic conditions. Several constituent strains, including E. faecium, L. plantarum and L. rhamnosus are known to produce GABA or stimulate its production, offering a mechanistic basis for this effect.
Of particular interest is the observed rise in L‑DOPA (3,4-dihydroxyphenylalanine, levodopa, precursor of dopamine) upon SymproveTM supplementation under high antibiotic‑induced dysbiosis. While L‑DOPA generated in the gut does not directly cross the blood-brain barrier, increased L-DOPA production and/or its reduced microbial decarboxylation could theoretically improve its systemic bioavailability during oral L‑DOPA therapy. Although speculative, such findings warrant further in vitro and in vivo exploration, especially given emerging evidence linking gut microbial activity to e.g. Parkinson’s disease treatment outcomes.
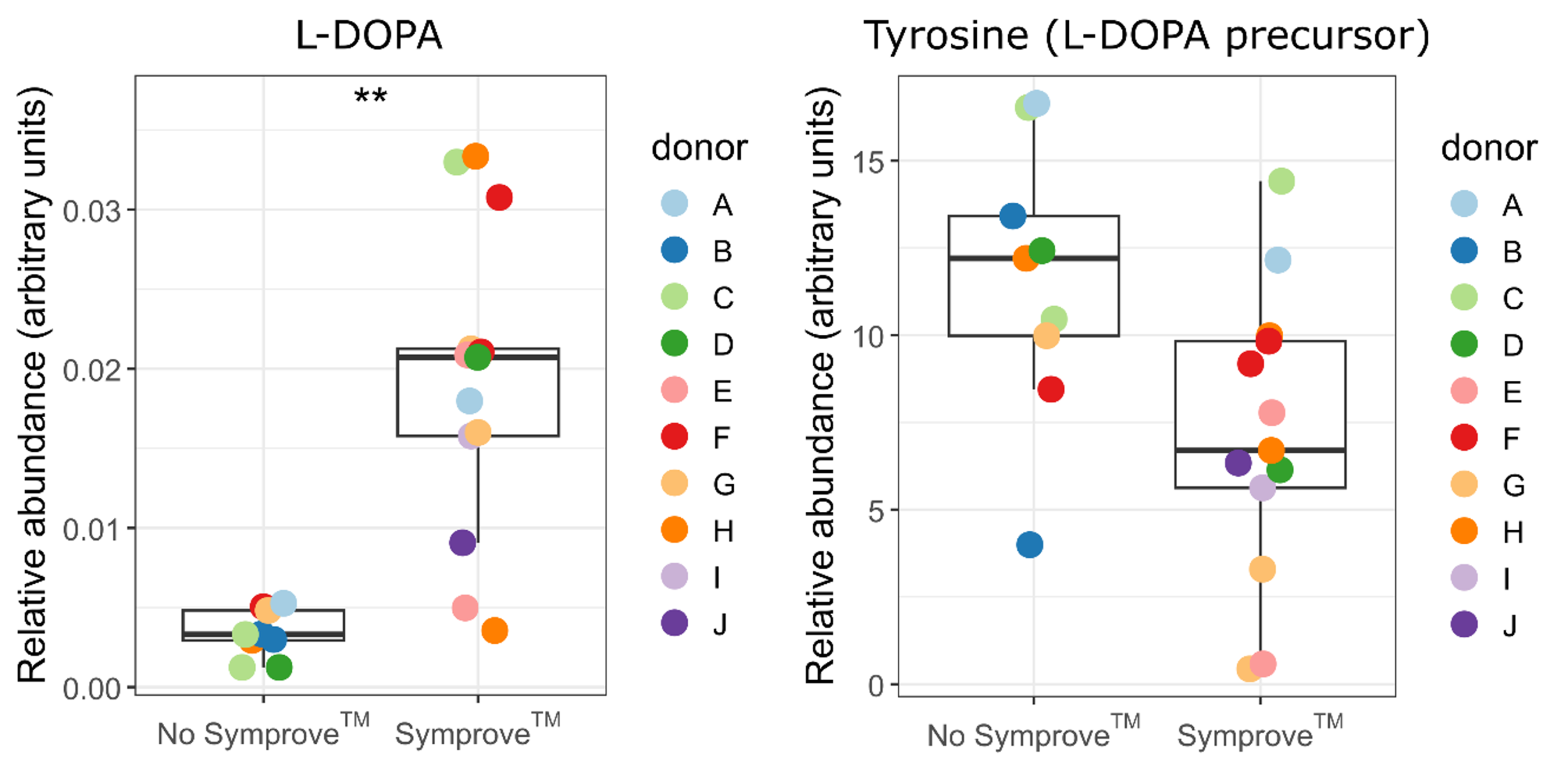
Impacts of Symprove™ on L-DOPA observed in the dysbiosis populations. UHPLC-HRMS profiling was used to quantify relative levels of metabolites from colonic fermentations collected after 24 h incubation with Symprove™ or untreated control (water) in the CoaP model using fecal samples from 10 healthy human donors. The low dysbiosis and high dysbiosis populations included the samples that were not infected with ETEC but were exposed to either low-dose or high-dose antibiotics, respectively, to induce dysbiosis. ETEC: enterotoxigenic Escherichia coli; L-DOPA: 2,4-dihydroxyphenylalanine; UHPLC-HRMS: ultra-high performance liquid chromatography-high-resolution mass spectrometry.
Conclusion: The Value of High‑Throughput Microbial Metabolism Platforms
This study highlights the ability of SymproveTM to modulate gut microbial metabolism across a remarkably diverse set of conditions, including severe dysbiosis. The combined use of Colon‑on‑a‑Plate®, LA‑REIMS , targeted SCFA analysis, and in-depth metabolomics using UHPLC-HRMS demonstrates the unique strength of our high‑throughput ex vivo colon microbiome platform to uncover metabolic mechanisms that remain invisible to conventional microbiome sequencing.
Beyond CoaP and metabolomics, we also offer powerful follow‑up platforms (including SHIME®, kinetic CoaP, mucosal simulations, upperGIT and host‑microbe interaction assays) that can further unravel the physiological implications of these findings and support translational development of next‑generation probiotics, postbiotics, and microbiome‑targeting therapeutics.
Want to know more? Read the full publication here!


