Recent Posts:
From compound lists to confident decisions: unlocking the full value of gut metabolomics
The gut microbiome is now widely recognized as a central driver of human health, influencing immunity, metabolic fitness, gut barrier integrity, the gut-brain axis and many more. Often the impact on health is induced by metabolites produced by the gut microbiome. As a result, metabolomics has become a key technology to assess how microbial activity translates into functional, host‑relevant outcomes. While metabolomics generates powerful datasets, data alone is not insight.
At ProDigest, we continuously push both our experimental models and our data analytics to help our clients move from complex datasets to biologically relevant decisions. Today, we take the next step in that journey.
Why gut metabolomics matters and why MetaKey® exists
Microbial composition tells you who is there. Metabolomics tells you what they are actually doing. Our MetaKey® platform provides a relevant view on gut microbial activity, more specifically:
- The gut metabolite profile: Quantification of 230 gut‑relevant metabolites, selected based on hundreds of scientific studies
- Coverage of key health functionalities (immunity, gut‑brain axis, metabolic health, gut barrier, inflammation, cardiac health) and disease concepts (e.g. IBD, obesity, diabetes, neurological and cardiovascular disorders)
- Strong biological breadth in terms of microbial vs host origin, chemical classes (>30) and metabolic pathways (>15)
- Flexible use through full profiles or focused panels, depending on your research question
MetaKey® allows researchers to explore how test products reshape gut metabolism and to identify useable endpoints for product evaluation, and even beyond the gut itself.
The challenge: when more data creates less clarity
As metabolomics datasets grow richer, interpretation becomes the real bottleneck. Traditionally, metabolomics analysis relied heavily on univariate statistics (assessing one metabolite at a time) and large collections of individual plots (each statistically correct, yet difficult to interpret as a whole). While this approach can be informative, it often:
- Lacks biological context
- Obscures overarching metabolic shifts
- Makes it hard to connect outcomes to mechanisms of action or study design
Manual pathway interpretation is possible but slow, inefficient and hard to scale. To solve this, ProDigest has upgraded MetaKey® with an integrated interpretation layer called SIMON® (Statistical Integration of Multi-Omics Networks).
SIMON® does not replace metabolomics, it unlocks its full value:
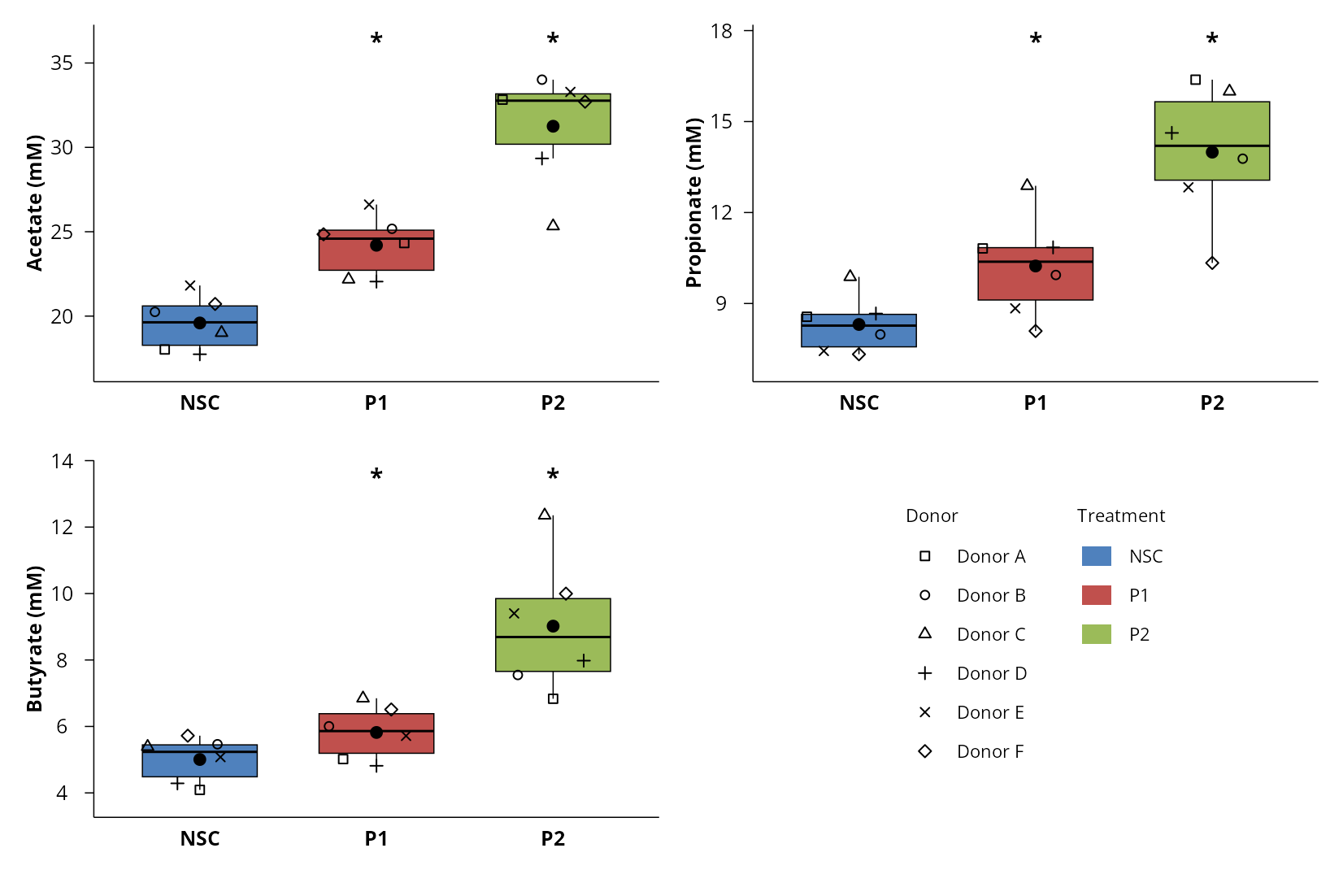
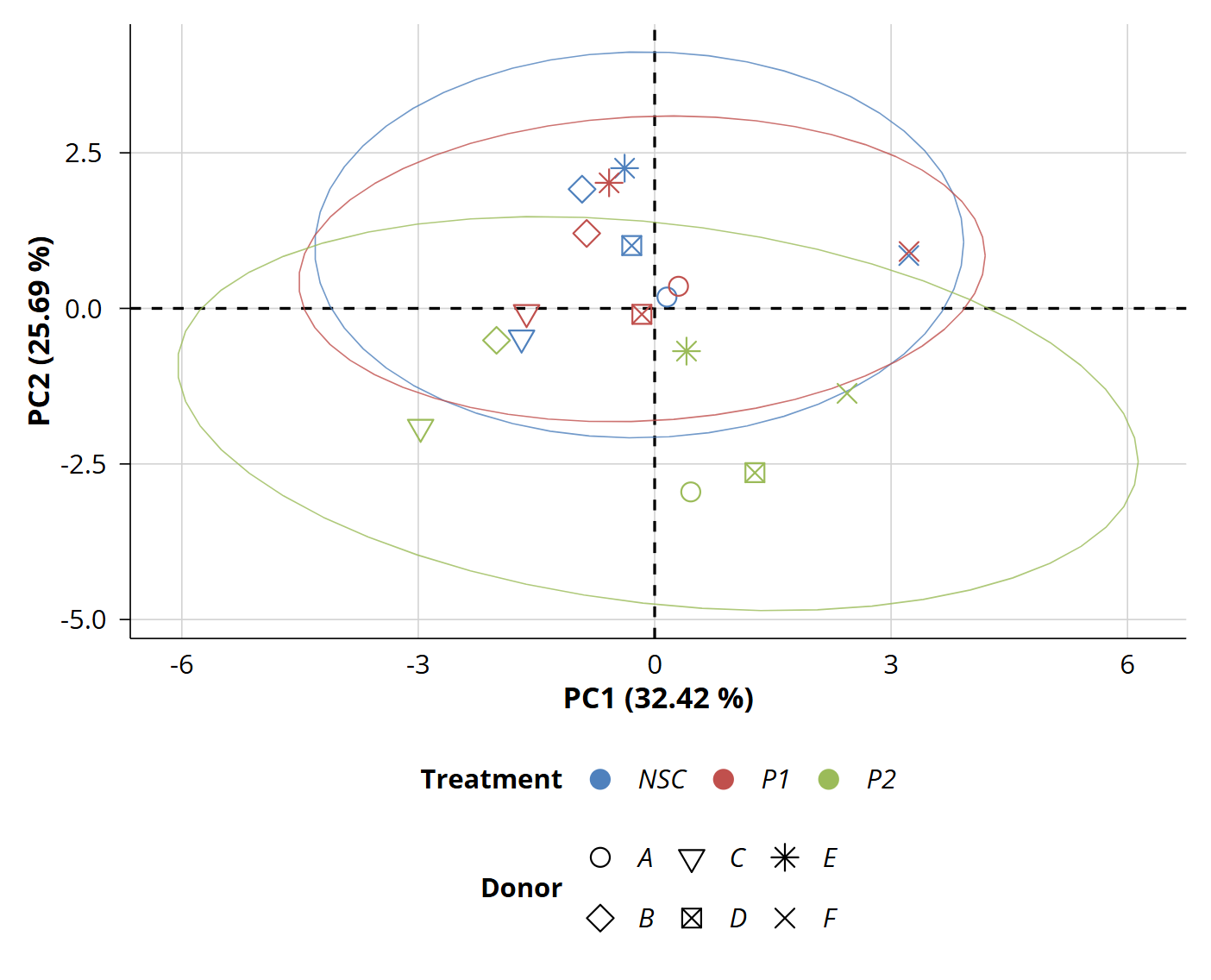
- Improved statistics and visualizations: optimized univariate statistics to address both targeted datasets, as well as more intuitive graphical outputs and clear overviews of impactful metabolites (see examples below).
- Gut‑specific pathway mapping: gut‑specific curation (focused on host-microbiota interactions), functional pathway interpretation (highlighting metabolic routing, e.g. saccharolytic vs. proteolytic fermentation) and exclusion of non‑gut‑relevant or misleading pathways.
Integrated biological interpretation: linking pathway level changes back to individual metabolites, connecting metabolomic outcomes to study design and experimental conditions, translating statistical output into mechanistic hypotheses.
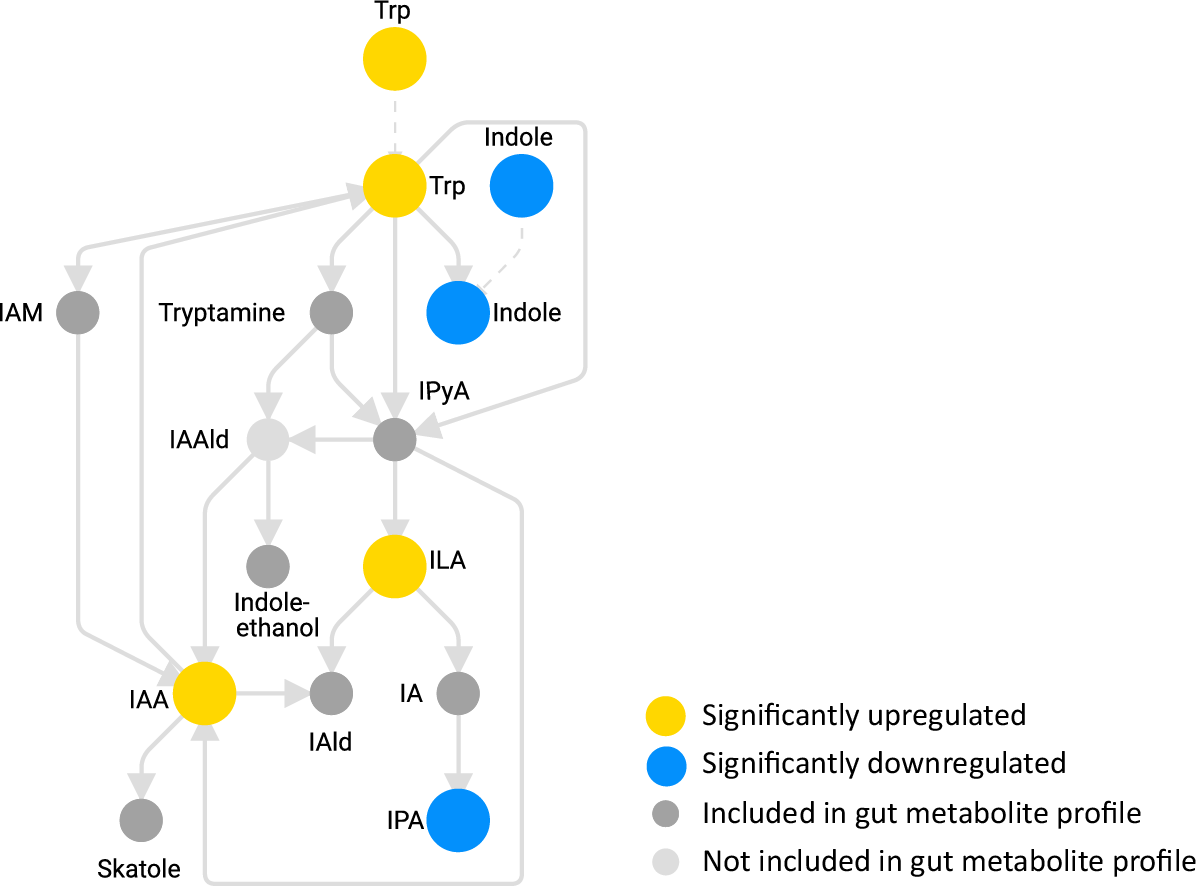
A reanalysis of our previously published SymproveTM study illustrates this clearly (see figure below): shifts in tryptophan metabolism highlighted increases in indole 3 lactic acid (ILA) and indole 3 acetic acid (IAA)-directly linking probiotic activity to microbiome driven immune and barrier effects.
Conclusion: beyond metabolomics, ready for multi‑omics
SIMON® is fully integrated with our established MetaKey® platform and is broadening its scope to support metabolomics and metagenomics integration. It has paved the way for our gut models towards multi‑omics analysis and enables mechanistic links between the microbiome and systemic physiology.
Want to know more? Let’s talk!


