
Recent Posts:
Synbiotic Strategies to Shape the Infant Gut Microbiome: Insights from Colon‑on‑a‑Plate®
Early life is a critical window for gut microbiome development with long‑term consequences for immune maturation, metabolic health and infection resistance. Breastfed infants typically harbor a microbiota dominated by Bifidobacterium, maintained by human milk oligosaccharides (HMOs) whereas formula‑fed infants often show a more heterogeneous and less bifidogenic profile. This has stimulated interest in adding prebiotics, probiotics and synbiotics to infant nutrition to better approximate the functional characteristics of the breastfed microbiome.
In this project, we have applied our Colon‑on‑a‑Plate® (CoaP) platform to evaluate the impact of two prebiotic fibers and one Bifidobacterium strain (on its own and in combination) on the gut microbiota of six healthy formula‑fed infants (<4 months). The study integrated fermentation readouts, 16S rRNA gene sequencing, flow cytometry and targeted UHPLC‑HRMS metabolomics to capture both community structure and functional metabolism under physiologically relevant colonic conditions.
Study Design: Multi-Parametric Assessment Across Six Infant Microbiomes
Cryopreserved fecal inocula from six formula‑fed infants were used as microbiome sources in our CoaP model. For each donor, eight conditions were tested over 24 hours:
- No prebiotics, no probiotic
- Non‑supplemented control (NSC)
- Prebiotics only
- F1 (single fiber)
- B (fiber blend)
- BF1 (combination of both fibers)
- Probiotic only
- + (Bifidobacterium sp.)
- Prebiotics combined with probiotic:
- F1+
- B+
- BF1+
The background medium represented the colonic environment of healthy, formula‑fed infants. Endpoints included:
- Fermentation parameters: pH, gas, SCFAs, branched SCFAs (BCFAs), lactate, ammonium
- Microbial composition and density: 16S rRNA gene Illumina sequencing + flow cytometry
- Gut metabolic profile: targeted UHPLC‑HRMS of 230 gut‑relevant metabolites (MetaKey® panel)
This design allowed for a robust comparison of treatment effects compared to the NSC group while explicitly capturing inter‑individual variability in response.
Fermentation Profiles: Synbiotic BF1+ as the Strongest Modulator
Across donors, the synbiotic BF1+ (blend of both prebiotics + Bifidobacterium) showed the most consistent and pronounced metabolic impact.
- BF1+ lowered colonic pH to values typical of healthy breastfed infants, driven by substantial increases in acetate and lactate.
- Blend‑containing conditions (B, BF1, B+, BF1+) significantly enhanced acetate production and shifted SCFA proportions toward an acetate‑dominated profile, closely resembling breastfed infant patterns.
- Propionate remained largely unchanged, while butyrate levels were low across conditions, as expected in young infants with limited complex fiber intake and fast transit time. In some blend/synbiotic conditions, butyrate was further reduced, likely due to pH‑mediated suppression of classical butyrate producers.
- BCFAs remained low (<0.5 mM) and were not differentially affected by treatment, reflecting a predominantly saccharolytic infant ecosystem.
- Ammonium was significantly reduced under BF1+, indicating a shift away from proteolytic fermentation toward saccharolytic metabolism (aligned with profiles typically observed in breastfed infants).
- Gas production did not increase under any treatment, including synbiotics, suggesting that these interventions are unlikely to promote gas-related discomfort and that they favor low‑gas producers such as Bifidobacterium.
Multivariate redundancy analysis (RDA) and discriminant analysis of principal components (DAPC) analyses confirmed that BF1+ sat at the extreme of a treatment gradient characterized by higher acetate/lactate and lower ammonium/BCFAs and consistently separated from NSC, F1 and probiotic‑only conditions.
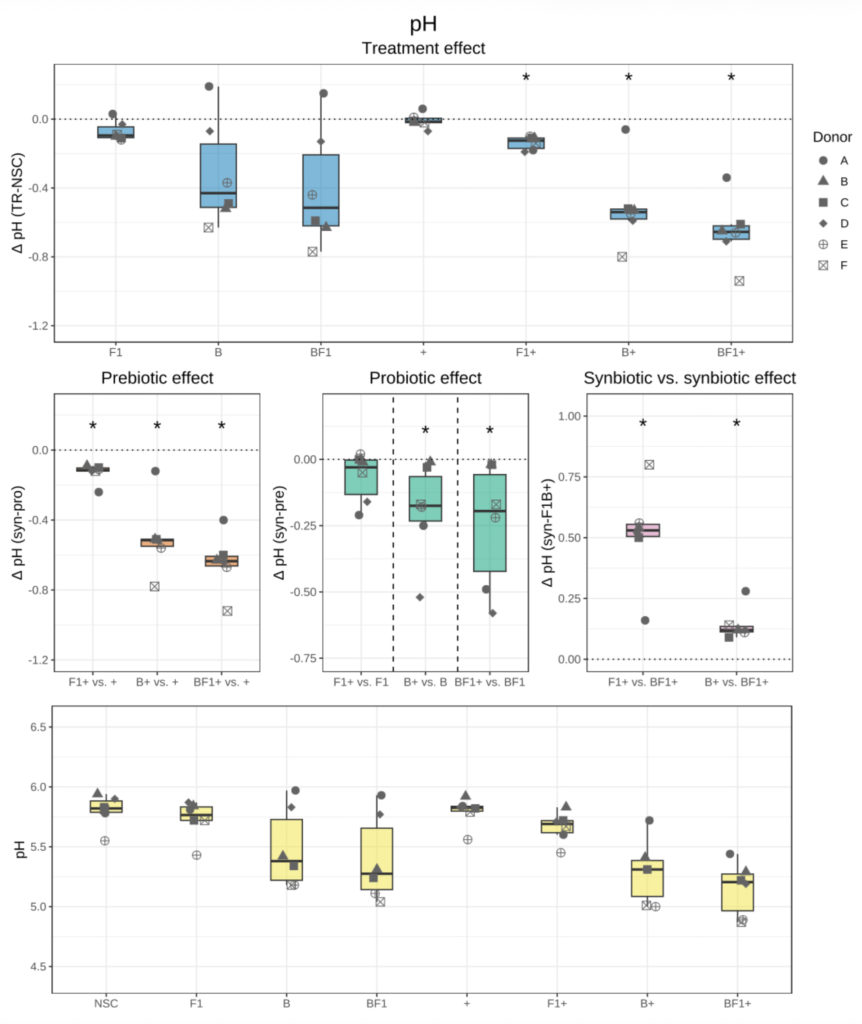
Box plot showing pH across the six healthy formula-fed donors (< 4 months) in the various conditions 24h after start of incubation (yellow). Difference in pH (Δ = Treatment – reference) between each treated condition and NSC (blue, treatment effect) at 24h across donors is given, as well as prebiotic (orange), probiotic (green), and synbiotic (pink) effects. The dotted line indicates no difference in pH for a given comparison. Statistical significance versus the compared condition (reference), based on paired Wilcoxon signed-rank tests, is indicated as follows: p < 0.05 (*), p < 0.01 (**), p < 0.001 (***), and p < 0.0001 (****).
Microbial Composition: Selective Bifidogenic Effects, Preserved Diversity
16S rRNA gene sequencing combined with flow cytometry showed that donor identity explained the majority of compositional variation, with three main baseline patterns: Bacillota/Proteobacteria‑, Bacteroidota‑ and Actinobacteriota‑(bifidobacteria‑) dominated communities.
Key findings:
- Alpha diversity (richness, evenness, Shannon, Gini‑Simpson) remained largely unchanged across treatments, indicating that the interventions were selective rather than disruptive to ecosystem structure.
- Beta diversity (PCA, PERMANOVA) showed strong donor clustering and minor but coherent treatment effects, with blend‑containing conditions, especially BF1+, forming distinct centroids in DAPC space
Differential abundance analysis (LEfSe, treeclimbR, MaAsLin2) highlighted:
- Enrichment of Bifidobacterium in all synbiotic conditions (F1+, B+, BF1+) and, more variably, in B and BF1 alone, supporting a clear bifidogenic effect of combining prebiotics with a Bifidobacterium strain.
- Reduction of Escherichia-Shigella across blend‑containing conditions and of Clostridioides (including difficile) in BF1+ only, suggesting suppression of clinically relevant pathobionts.
At amplicon sequence variant (ASV) level, a Bifidobacterium variant corresponding to the supplemented strain was consistently enriched in probiotic and synbiotic conditions and showed best engraftment in donors with low endogenous Bifidobacterium, while “priority effects” limited engraftment in already bifidobacteria‑rich microbiomes.
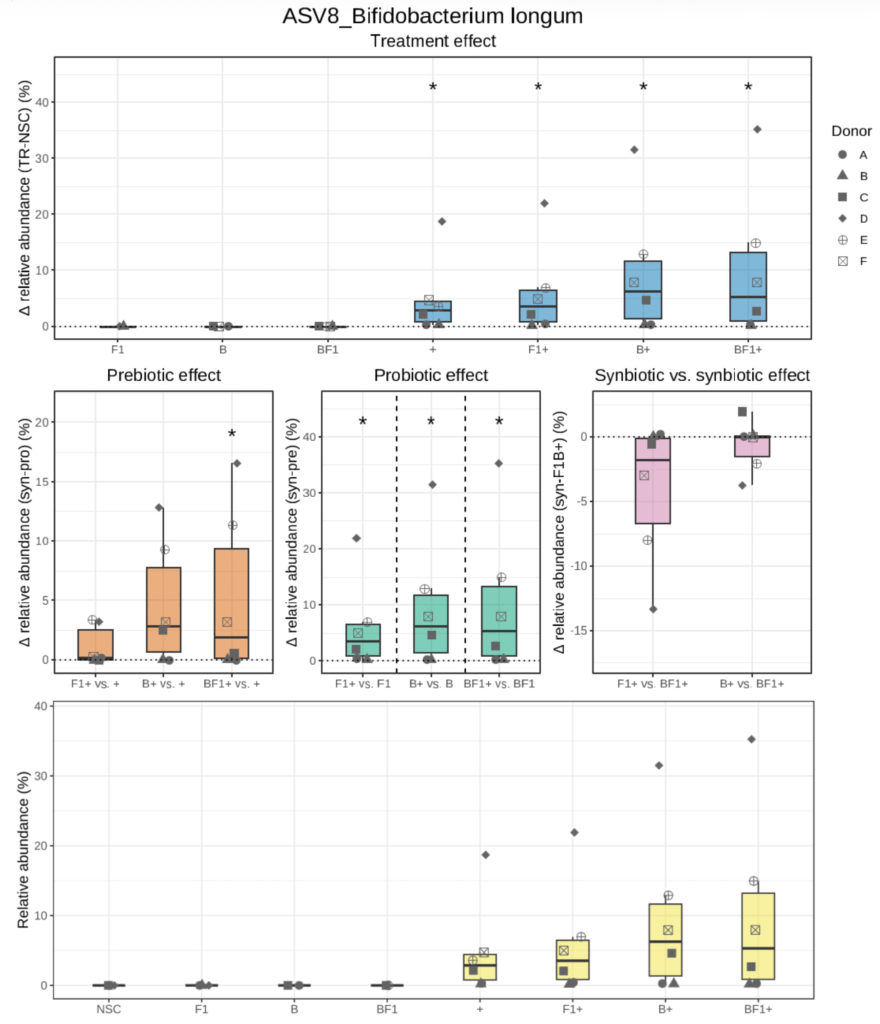
Box plot showing AV8 relative abundance across the six healthy formula-fed donors (< 4 months) at 24h (yellow), and difference (Δ = Treatment – reference) between each treated condition and NSC (blue, treatment effect), as well as prebiotic (orange), probiotic (green), and synbiotic (pink) effects, across donors at 24h. The dotted line indicates no difference for a given comparison. Statistical significance versus the compared condition (reference), based on MaAsLin2, treeclimbR, and LefSe analysis.
Metabolomics: Pathway-Level Support for Synbiotic Mechanisms
Targeted UHPLC‑HRMS profiling quantified 112 metabolic features above LOQ. Of these, 35 were significantly altered in at least one treatment vs NSC, with the largest changes observed for BF1, B+ and BF1+. Pathway analysis (MetaboAnalyst) identified enrichment of routes including branched‑chain amino‑acid metabolism, TCA cycle, glyoxylate and dicarboxylate metabolism and alanine/aspartate/glutamate metabolism.
Particularly interesting observations include:
- Increased GABA levels in several blend/synbiotic conditions, suggesting potential gut-brain axis relevance.
- Modulation of asparagine, citric acid, succinic acid and pyruvic acid, pointing to shifts in energy and nitrogen metabolism.
- Multi‑omics correlations revealed strong positive associations between Bifidobacterium and acetate, as well as metabolites like N‑acetylglucosamine and pyridoxine (vitamin B6) in synbiotic conditions, supporting a mechanistic link between bifidogenic shifts and beneficial metabolite production.
Preservation of Original Microbiome
The study showed that our CoaP model preserves each infant donor’s original microbiome structure throughout incubation, especially in the NSC condition, providing:
- High ecological fidelity, keeping donor‑specific communities intact rather than drifting toward a generic in vitro profile.
- A reliable baseline, so treatment effects can be confidently attributed to the ingredients, not to model artifacts.
- Retention of natural inter‑individual variation, with strong donor identity maintained across the study.
- Validated model conditions, as the medium supports microbial growth without distorting community composition, confirming physiological relevance.
Conclusion: Colon‑on‑a‑plate® as a Platform for Infant Synbiotic Discovery
This study demonstrates how combining two complementary prebiotics with a Bifidobacterium strain can drive the infant microbiome toward a more saccharolytic, bifidobacteria‑rich, breastfed‑like functional state, without disrupting overall diversity or increasing gas. The Colon‑on‑a‑Plate® platform was instrumental in generating these insights across multiple infant microbiomes in a controlled, high‑content manner.
Beyond CoaP and metabolomics, we also offer powerful follow‑up platforms (including SHIME®, host‑microbe interaction assays, static and dynamic digestion) that can further unravel the physiological implications of these findings and support translational development of next‑generation probiotics, prebiotics, postbiotics, and microbiome‑targeting therapeutics.
Want to find out more about what we can do? https://prodigest.eu/blogs


